|
8/22/2023 0 Comments Macvector aligning rnaseq readshashtype|-t = select hash table data structure to use: hashbits|-b = use n bit hashes (for n's of 1 to WORDSIZE. quickhash|-q = specify a hashing function:ġ - don't pack bits to make hash (use first word only)Ģ - naively use the first hashbits worth of patternģ - adaptivevely find a good hash (default) name|-n = alignment name (default: test) directory|-d = output directory (default: output) Using the format automatically generates a $ docker run -rm -it biocontainers/murasaki:v1.68.6-8b1- deb_cv1 murasaki -h > docker run -rm -it biocontainers/murasaki:v1.68.6-8b1- deb_cv1 murasaki -h MUMmer4: A fast and versatile genome alignment system Stefan Kurtz*, Adam Phillippy†, Arthur L Delcher†, Michael Smoot†‡, Martin Shumway†, Corina Antonescu† and Steven L Salzberg† 3、 Versatile and open software for comparing large genomes 2、 Fast algorithms for large-scale genome alignment and comparison.ĭelcher AL1, Phillippy A, Carlton J, Salzberg SL. SnapGene software (from GSL Biotech available at ) 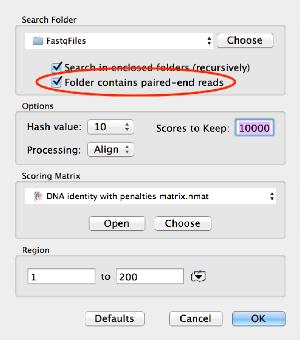
SnapGene Viewer | Molecular Biology Software SnapGene Software Tutorial Videos | Aligning to a Reference DNA Sequence SnapGene | Software for everyday molecular biology
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
